KNIME Workflows
As our goal is to make CHEESE Search a fully customizable tool that can be easily integrated into the research pipeline, we also support KNIME workflows, which are frequently used by computational chemists.
Searching by CSV
For a simple search using CSV files, we provide a template KNIME Component that processes a list of SMILES strings and returns the results as a CSV file.
CHEESE Search API component allows the user to define the list of query molecules and the similarity search parameters, including the number of neighboring molecules to retrieve, the type of similarity metric, and the speed/accuracy level, along with other options exposed by the CHEESE Search API. An example KNIME workflow using the CHEESE Search API component is shown below:
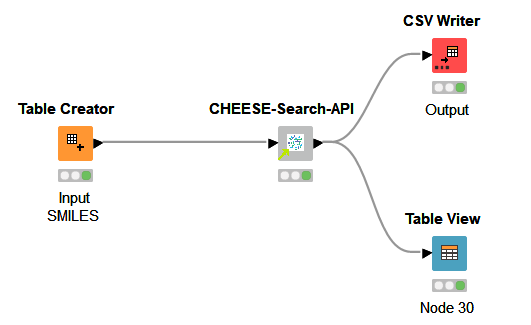
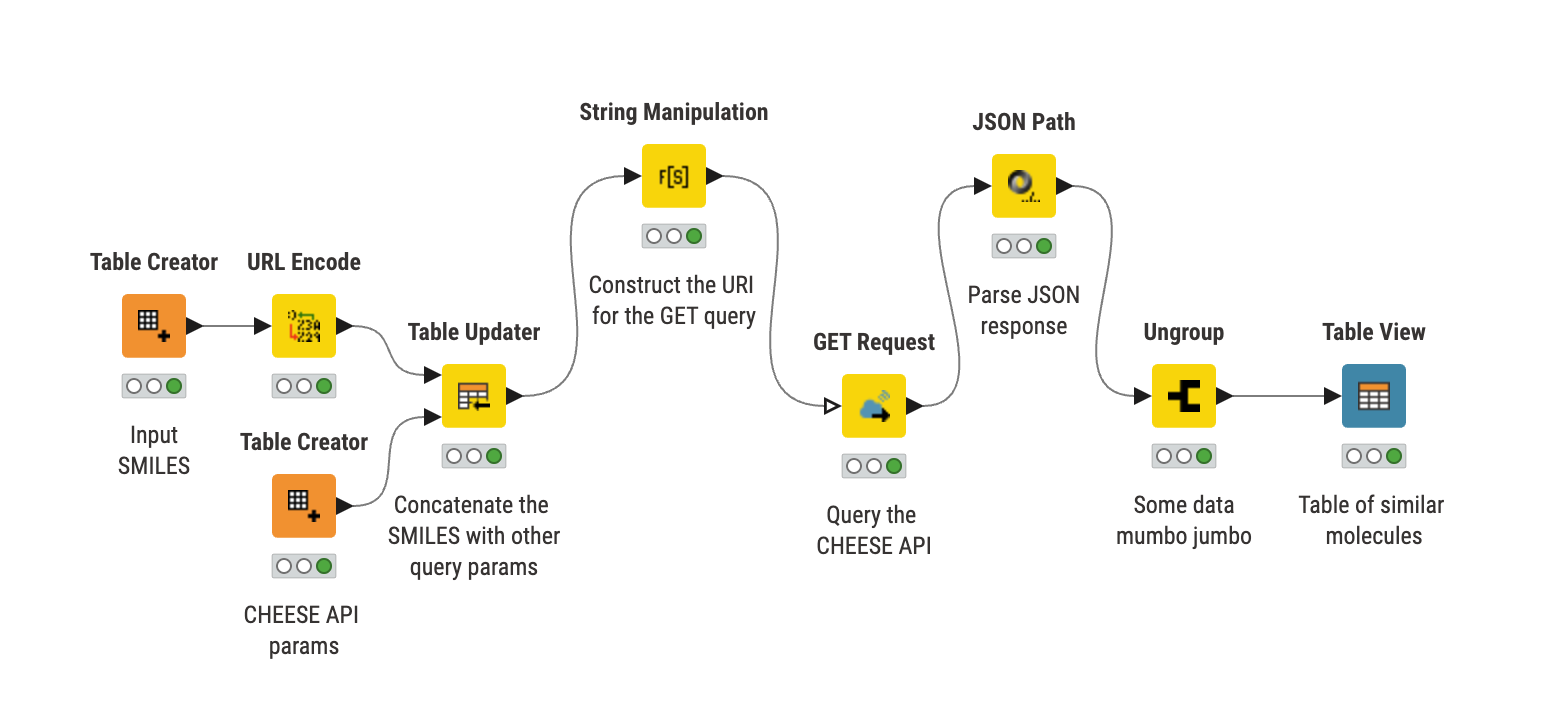
In more detail, the input SMILES are sent to the CHEESE search via an API request. The resulting JSON response is then processed into a list of cosine-similar molecules displayed in the output table.
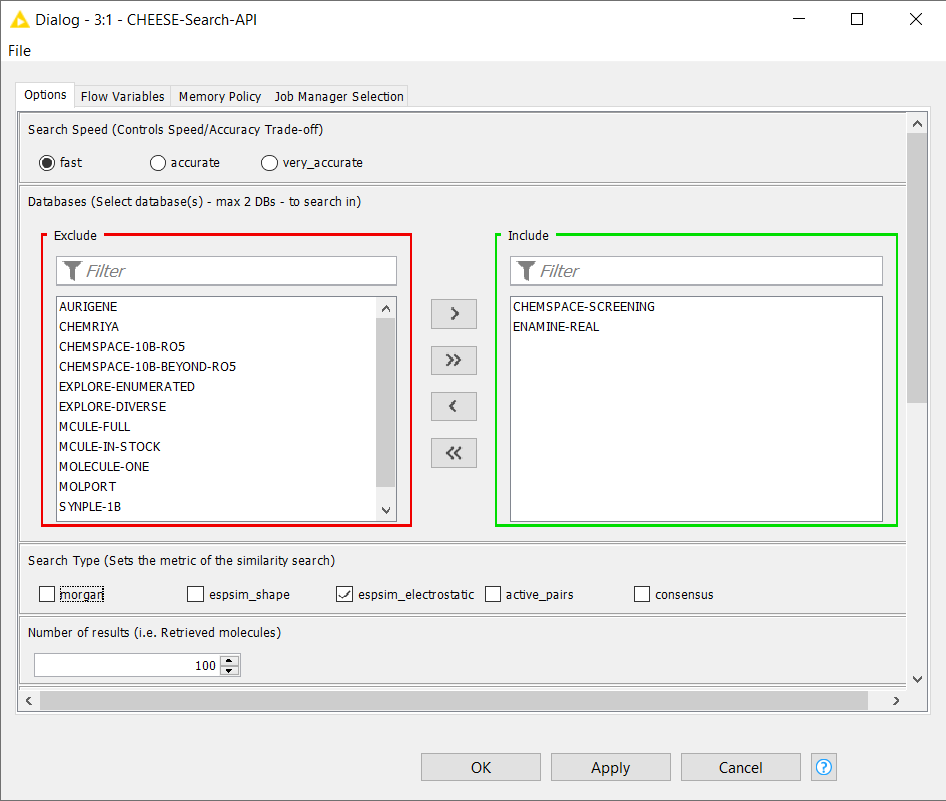
Within the CHEESE Search API component, users can select different databases, define the search accuracy, choose the search type (Morgan fingerprints, electrostatic or shape similarity, active pairs, or consensus), and specify the number of resulting molecules.
!
You can download the complete KNIME workspace here.
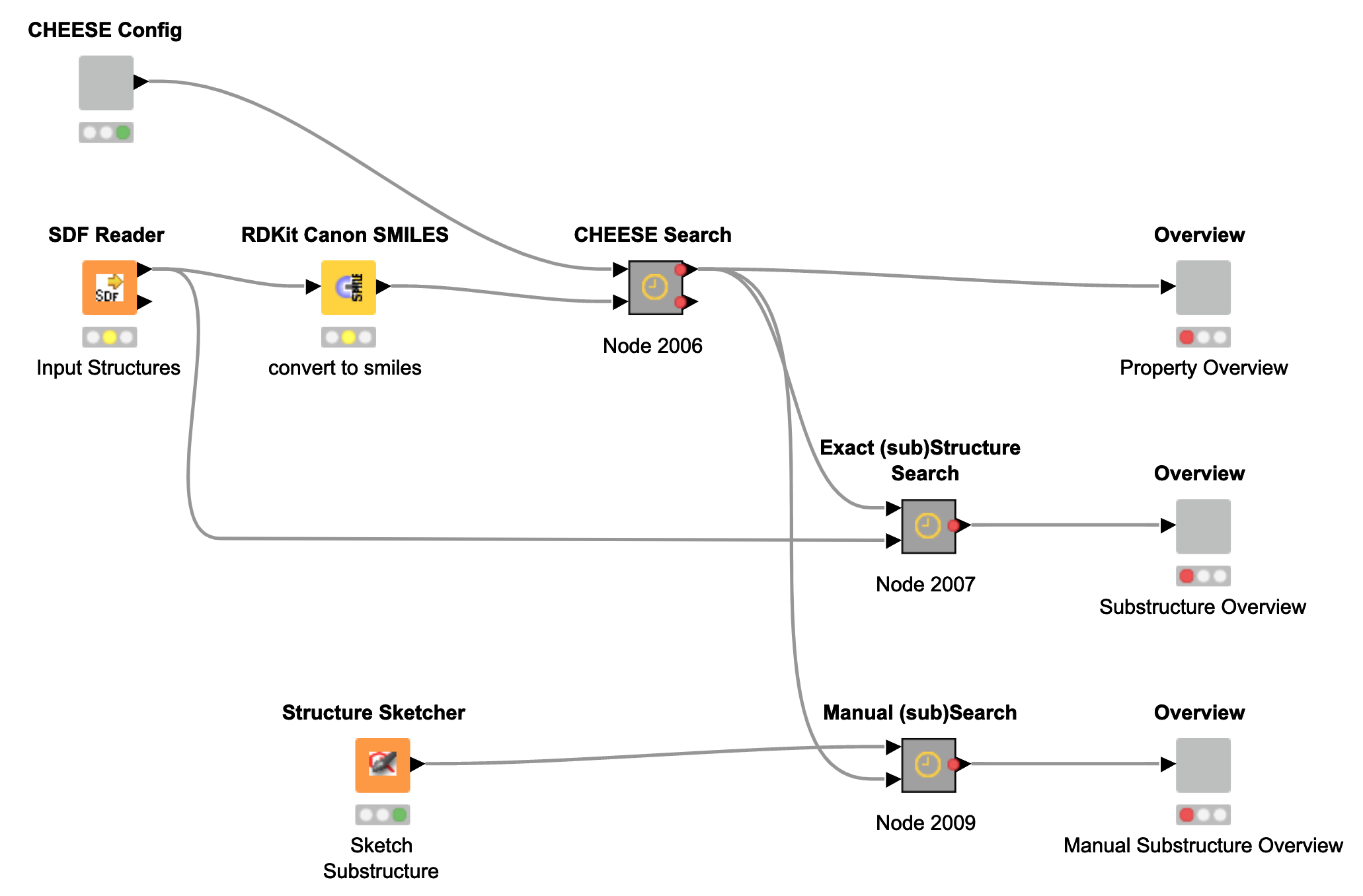
Searching by SDF (Contributed by Jan Skácel)
In addition to CSV-based searches, we also provide a KNIME workflow for searches using SDF files as input. For this purpose, the RDKit KNIME extension is required. Using the Search by SDF component, users can also draw molecular structures directly via the Structure Sketcher node.
The Search by SDF component can be downloaded here.